To access PalmXplore homepage, go to:
http://palmxplore.mpob.gov.my/
Alternatively, PalmXplore can be accessed via a direct link from genomsawit website (http://genomsawit.mpob.gov.my/)

| Section | Description |
| A | Quick Search Gene and/or gene model may be searched by gene id, scaffold id, keywords, specific gene and/or location. Clickable examples with correct format are provided. Search term examples include: |
| i. Location: p5_sc*****:******-****** The format for location search starts with the oil palm P5 scaffold id (example: p5_sc[00001 to 40360]), a colon (:), start and end location with a (-) separator. Search term example: p5_sc00060:424800-446800
|
|
| ii. Gene Id: p5.00_sc*****_p**** "p5.00" indicates the genome version, which is P5 build. "sc*****" indicates scaffold id. "p****", the digits after the "p" are auto-incrementally-generated by positions in the genome. Search term example: p5.00_sc00060_p0019
|
|
| iii. Scaffold ID: p5_sc***** "p5" indicates the oil palm genome version, which is P5 build. "sc*****" indicates scaffold id. Search term example: p5_sc00060
|
|
| iv. Specific Gene(s): a) Intronless gene: EgIG_*
"Eg" indicates species of oil palm (Elaeis guineensis). "IG" identifies the gene as intronless. Digit after "EgIG_" indicates intronless gene number. Search term example: EgIG_1 OR intronless gene
b) Fatty acid biosynthesis gene: Eg*****_*
"Eg" indicates species of oil palm (Elaeis guineensis). Letters after "Eg" indicates fatty acid gene name (e.g. ACC, CT, BCCP, BC, FABD, FABH, FABB, FABF,FABG, FABZ, FABI, FAB2, FAD2, FAD3, FATB or FATA). Digit after "_" indicates gene number. Search term example: EgACC_1 OR FAB genes
c) Resistance gene: Eg_rgh_***_*
"Eg" indicates species of oil palm (Elaeis guineensis). "rgh" identifies the gene as a resistance gene. Letters after "Eg_rgh_" indicates class of the resistance gene (e.g. cnl, kinase, mlo, rlk, rlp or others). Digit after "Eg_rgh_***_" indicates resistance gene number. Search term example: Eg_rgh_cnl_1 OR Resistance genes
d) Shell gene: Search term example: SHELL OR Shell gene
e) Virescens gene: Search term example: Eg_VIR_MYB113 OR VIRESCENS OR Virescens gene
f) Mantled gene: Search term example: EgDEF1 OR MANTLED OR Mantled gene
|
|
| v. Other search terms: a) Enzyme Code (EC): Search term example: EC:3.5.1.98
b) GO ID: Search term example: GO:0016810
c) PFAM ID: Search term example: PF00850
|
|
| B | Please cite us! Guide on how to cite PalmXplore system and its underlying publication(s). |
| C | Quick Links |
| i. Advanced Search tool: Multiple options to select specific data types and parameters to formulate queries. |
|
| ii. CDS Browser: Thematic browser that lists all of the coding sequence (CDS) identifiers associated with the predicted oil palm genes. |
|
| iii. SCAFFOLD Browser: Thematic browser that lists all of the genomic scaffolds from E. guineensis P5 build. |
|
| iv. MYPalmViewer: Explore and navigate the chromosomes of oil palm, it's annotations and associated data tracks. |
|
| v. BLAST tool: The integrated BLAST tool provides BLASTN, BLASTX, TBLASTX and TBLASTN programs to match a query sequence to oil palm sequences. |
|
| vi. Download : List of data records of genome assemblies and gene annotation. |
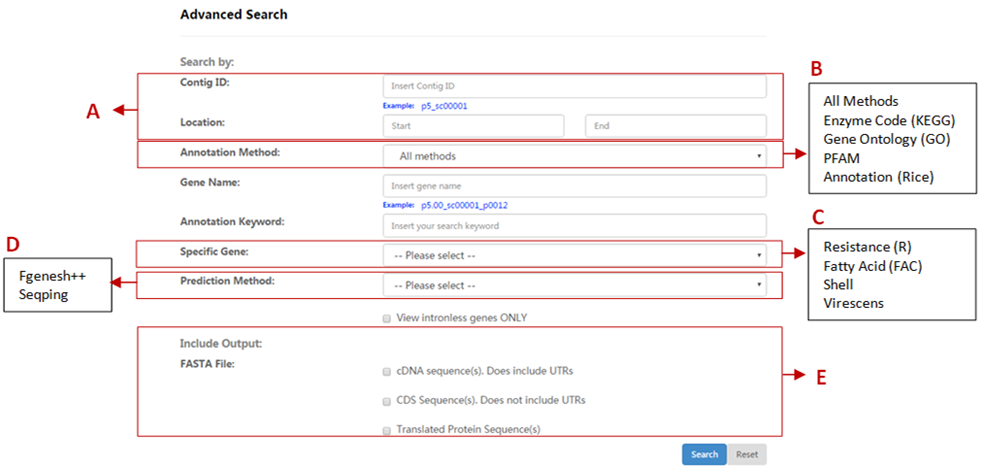
| Section | Descriptions |
| A | Scaffold ID and Location
Search genes within a genomic scaffold and location by providing Scaffold ID and/or Start and End coordinates. |
| B | Annotation Method
Genes can be searched based on Enzyme Code (EC), GO, PFAM and rice annotation from BLAST. |
| i. All methods Perform a cross search from Enzyme Code, GO and PFAM annotation results. |
|
| ii. Enzyme Code (EC) Upon selecting this method, Enzyme Code (EC) input field is enabled. Search term example: EC:3.5.1.98
|
|
| iii. Gene Ontology (GO) Upon selecting this method, GO ID/GO term input field is enabled. Search term example: GO:0016810
|
|
| iv. PFAM Upon selecting this method, PFAM ID input field is enabled. Search term example: PF00850
|
|
| v. Annotation from BLAST Filter search according to the rice annotation from BLAST. |
|
| C | Specific Gene Restrict gene searching to specific genes, such as Resistance, Fatty Acid Biosynthesis, Shell, Virescens and Intronless. |
| D | Prediction Method Restrict gene searching according to prediction method; Seqping or Fgenesh++. |
| E | Include Fasta File Include FASTA-formatted sequence file(s) (view/download) for: a. cDNA sequence(s), which include UTRs b. CDS Sequence(s), which does not include UTRs c. Translated Protein Sequence(s) |
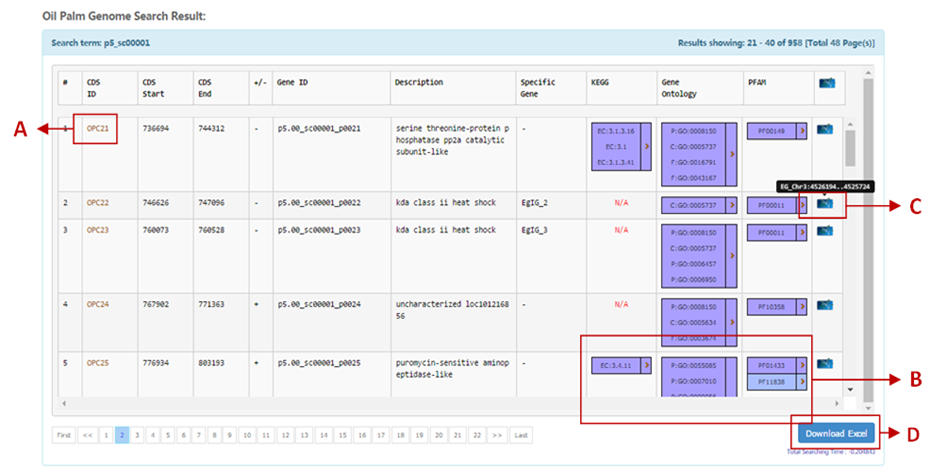
| Section | Descriptions |
| A | CDS Details View details of the selected CDS 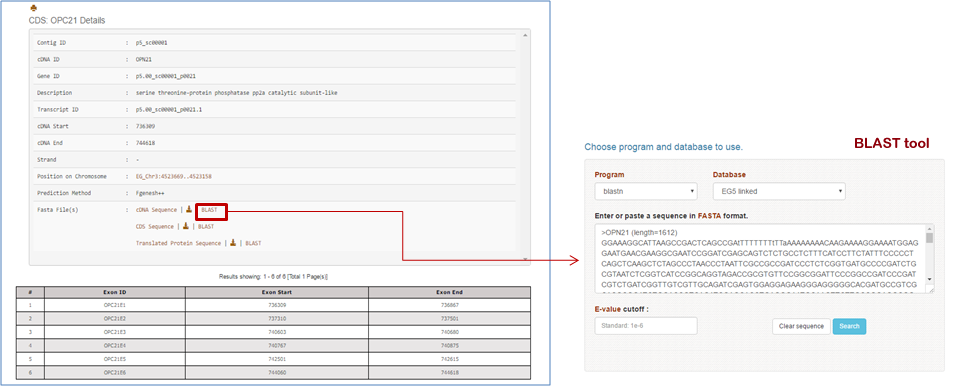
View BLAST tool help page |
| B | Enzyme Code (EC), GO and PFAM View details of the selected CDS associated with functional annotation from Gene Ontology (GO), PFAM and KEGG databases (including links to the external databases). |
| C | MyPalmViewer (GBrowse) Click on the View MyPalmViewer help page. |
| D | Download Click "Download Excel" button to download the populated data from the query in MS Excel format. Downloading a large retrieval may take several minutes. |



